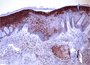
Microscopic examination of tissue morphology is an important initial step in the phenotypic analysis of normal and diseased cutaneous tissue. Optimal histologic skin preparations are central to this analysis. The Core provides protocols, training, for fixation, decalcification (if needed), trimming of tissue, and microtomy. Investigators provide fixed, trimmed tissue samples in labeled cassettes to the Core for processing, embedding into paraffin, sectioning onto glass slides, and H&E staining. Select special histochemical stains are also available.
In addition to standard samples of mouse, human, and pig skin, the core has expertise with a variety of other skin specimens that include xenografts, ex vivo organ culture of full thickness skin and microdissected scalp hair follicles, in vitro / in vivo organotypic constructs, reconstituted hair follicle patch/chamber constructs, tumors, and skin wounds. The core can also process, such as embryos, nail, teeth, tongue, oral mucosa, palate, mammary gland, and various internal organs. However, a consultation with the Core is required before a non-routine request can be submitted.
Normal and diseased archived human skin tissue is invaluable for translational studies. The CPAT Core has access to an extensive and well-characterized repository of clinical samples that are formalin-fixed and embedded in paraffin wax. SBDRC members may request slides to be sectioned from relevant clinical samples. Members request samples using the online form, and consultation with the CPAT Core staff, paraffin blocks from selected clinical cases we request archival samples for the project. Once the samples arrive in the lab, investigators are notified and then submit an online order for slides and histochemical stains. Additional services, such as LCM or immunostaining, are also available for use on clinical slides generated by the Core.
As FFPE samples are sometimes not compatible with experimental approaches, the CPAT core offers cryo sample services including training for embedding in cryopreservation media, sectioning of frozen samples, and access to a cryostat for frozen slide preparation by investigators. The Core also offers limited cryosection services for pilot studies and small projects.
In addition to morphology, analysis of molecular changes are critical for a mechanistic understanding of homeostatic and pathologic cutaneous biology. Analysis of protein expression and signal transduction is routinely carried out by the CPAT Core using immunohistochemistry and immunofluorescence on paraffin and frozen sections of skin.
Laser Capture Microdissection (LCM) enables isolation of homogeneous subsets of cells from the same tissue section, for example adjacent areas of epidermis that contain squamous cell carcinoma and normal epidermis. From the isolated cells, RNA and DNA is purified for use in downstream assays such as QPCR, next-generation-sequencing, and microarrays. The CPAT Core is home to a state-of-the-art Leica LMD 7000 microscope. After an initial consultation with CPAT Core personnel to discuss goals and experimental design, the Core will provide training on the LMD and for added flexibility, investigators have access to protocols, training, and all necessary equipment to cut and stain their samples. The Core also offers slide preparation services – FFPE or Frozen – investigators-provided polyethylene napthalate (PEN)-membrane slides, with their tissue sections under nuclease-free conditions.
Localization of specific mRNAs in tissue is an important part of understanding changes identified at the transcriptomic level. Compared to traditional methods for in situ hybridization, RNAscope technology provides superior sensitivity and specificity. Chromogenic and fluorescent readout options allow for flexibility in experimental design, with ability for duplex and triplex assays, respectively. Ease of use is enabled by validated reagents and protocols, as well as available probes to over 13,000 mRNA transcripts. The CPAT core has dedicated equipment and bench space for RNAscope assays. Once a member is trained, they can schedule the bench using our online calendar. addition, this service is valuable to SBDRC members who want to try the technology before investing in the equipment for their own labs. Limited regents, including OPAL dyes, may be available thorugh the core.
Skin barrier function is a crucial role of the epidermis, and impaired barrier is associated with a range of pathological conditions. Clinical diseases vary in severity from common disorders such as atopic dermatitis, to more rare conditions such ichthyosis which in some forms can be fatal. Assessing the state of the skin barrier is a critical tool for researchers investigating normal and aberrant epidermal differentiation. The CPAT Core of the Penn SBDRC offers services to determine skin barrier function by evaluating 1) corneometry to assess skin water content by the capacitance method. 2) skin pH measurements using the Skin-pH-Meter with a PH 905 probe and 3) transepidermal water loss (TEWL) by diffusion using a Tewameter TM300. TEWL and pH measurements can be made on haired and non-haired human or animal skin, though corneometry requires nonhaired, sparsely haired or shaved skin. Questions regarding these services can be sent to Dr. Charles Bradley (cbradle2@vet.upenn.edu) an expert in skin barrier function, located at Penn Vet.
Coming Soon.
Coming Soon.
 Pricing
Pricing Protocols
Protocols Tissues
Tissues Histochemical Stains
Histochemical Stains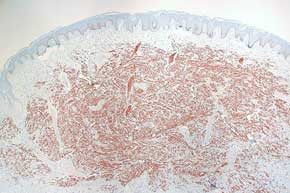 Immunostaining
Immunostaining RNAscope in situ hybridization
RNAscope in situ hybridization FAQ
FAQ Publications
Publications